SPANDx: a genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets, BMC Research Notes

VDAP-GUI: a user-friendly pipeline for variant discovery and annotation of raw next-generation sequencing data

Whole genome sequencing reveals extensive community-level transmission of group A Streptococcus in remote communities, Epidemiology & Infection

Using whole-genome sequencing and a pentaplex real-time PCR to characterize third-generation cephalosporin-resistant Enterobacteriaceae from Southeast Queensland, Australia
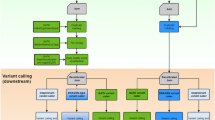
Comparative analysis of whole-genome sequencing pipelines to minimize false negative findings
Tracing the environmental footprint of the Burkholderia pseudomallei lipopolysaccharide genotypes in the tropical “Top End” of the Northern Territory, Australia

Generalizable characteristics of false-positive bacterial variant calls
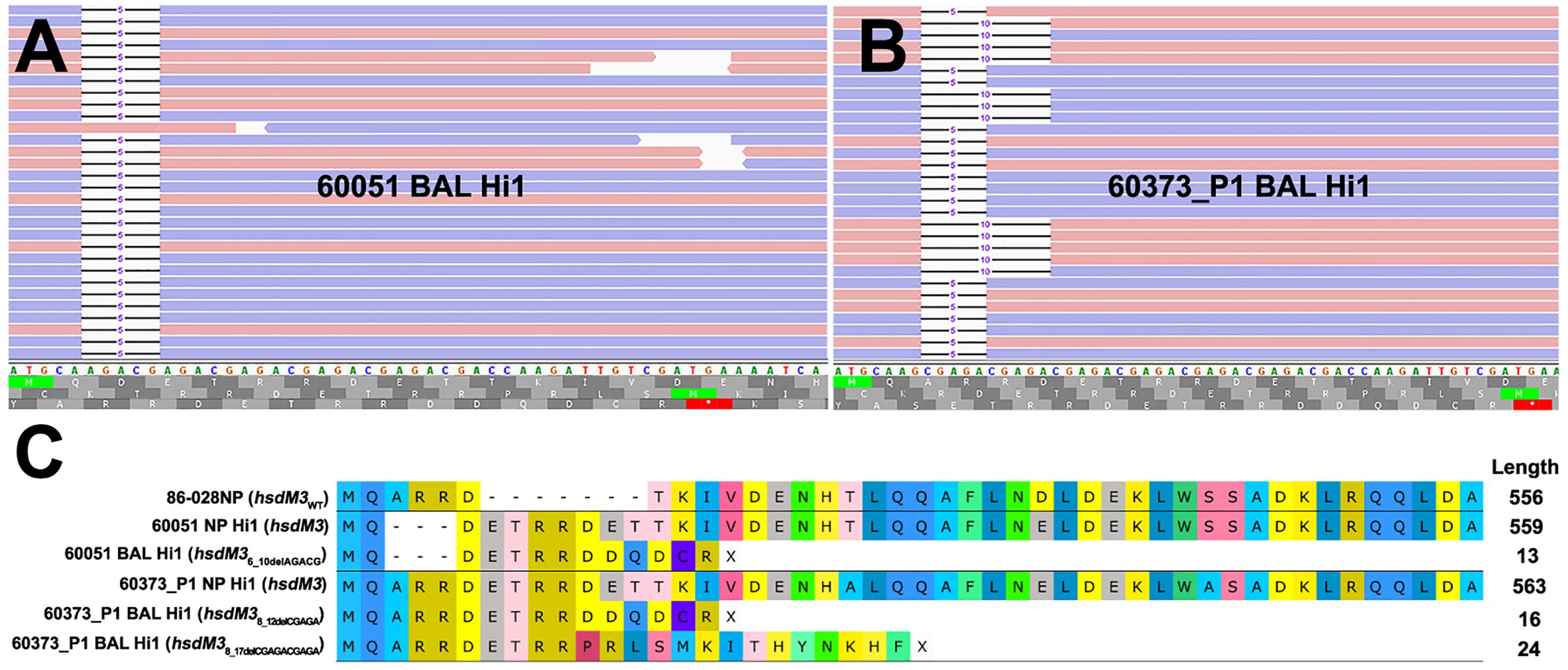
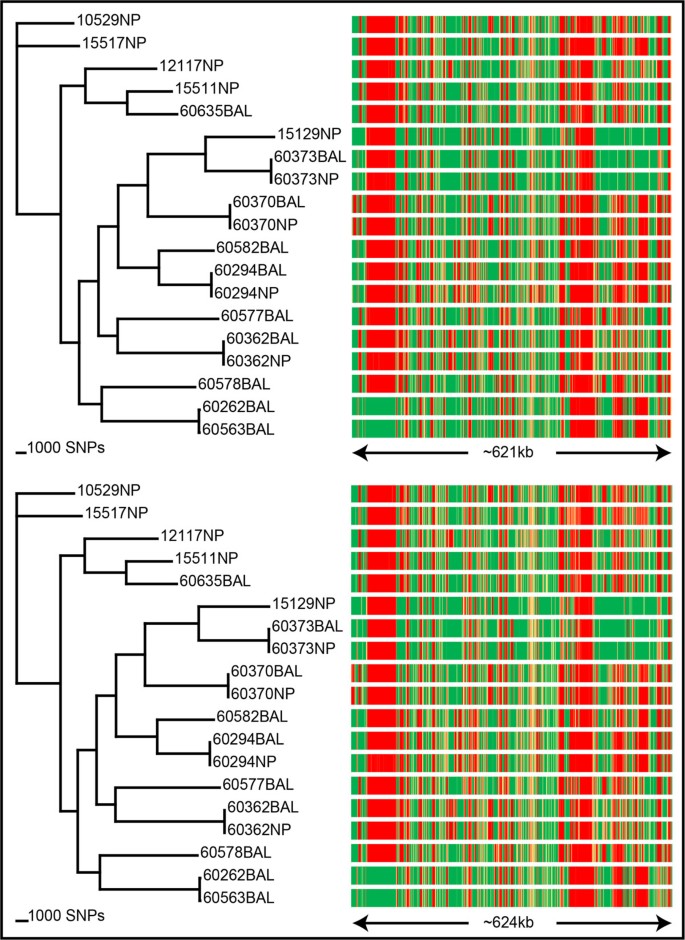
Frontiers Molecular Signatures of Non-typeable Haemophilus influenzae Lung Adaptation in Pediatric Chronic Lung Disease
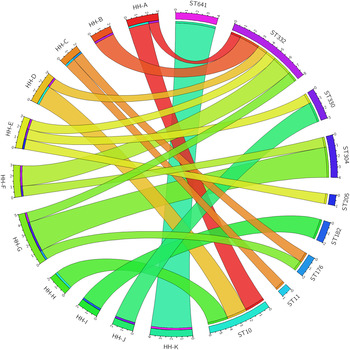
Whole genome sequencing reveals extensive community-level transmission of group A Streptococcus in remote communities, Epidemiology & Infection

The relationship between the alignment time and the FASTQ file size.

Computational methods for chromosome-scale haplotype reconstruction, Genome Biology

Comparative genomics confirms a rare melioidosis human-to-human transmission event and reveals incorrect phylogenomic reconstruction due to polyclonality
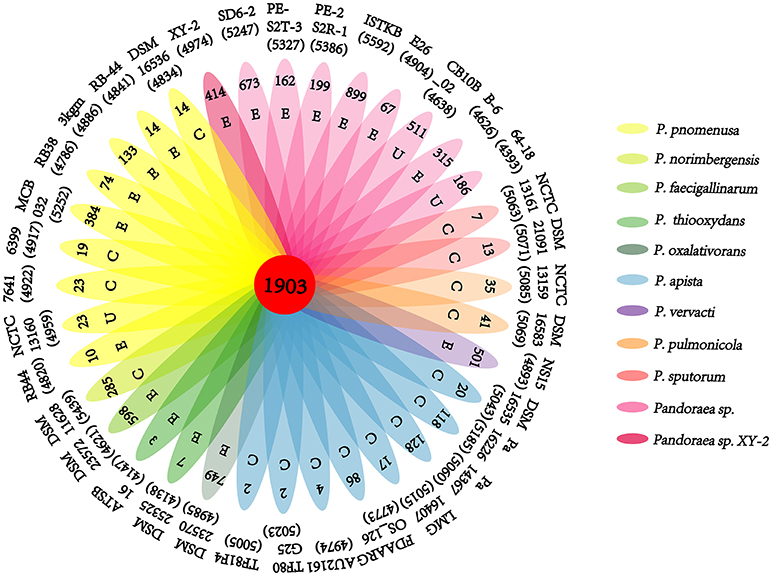
Frontiers Whole Genome Sequencing and Comparative Genomics Analyses of Pandoraea sp. XY-2, a New Species Capable of Biodegrade Tetracycline
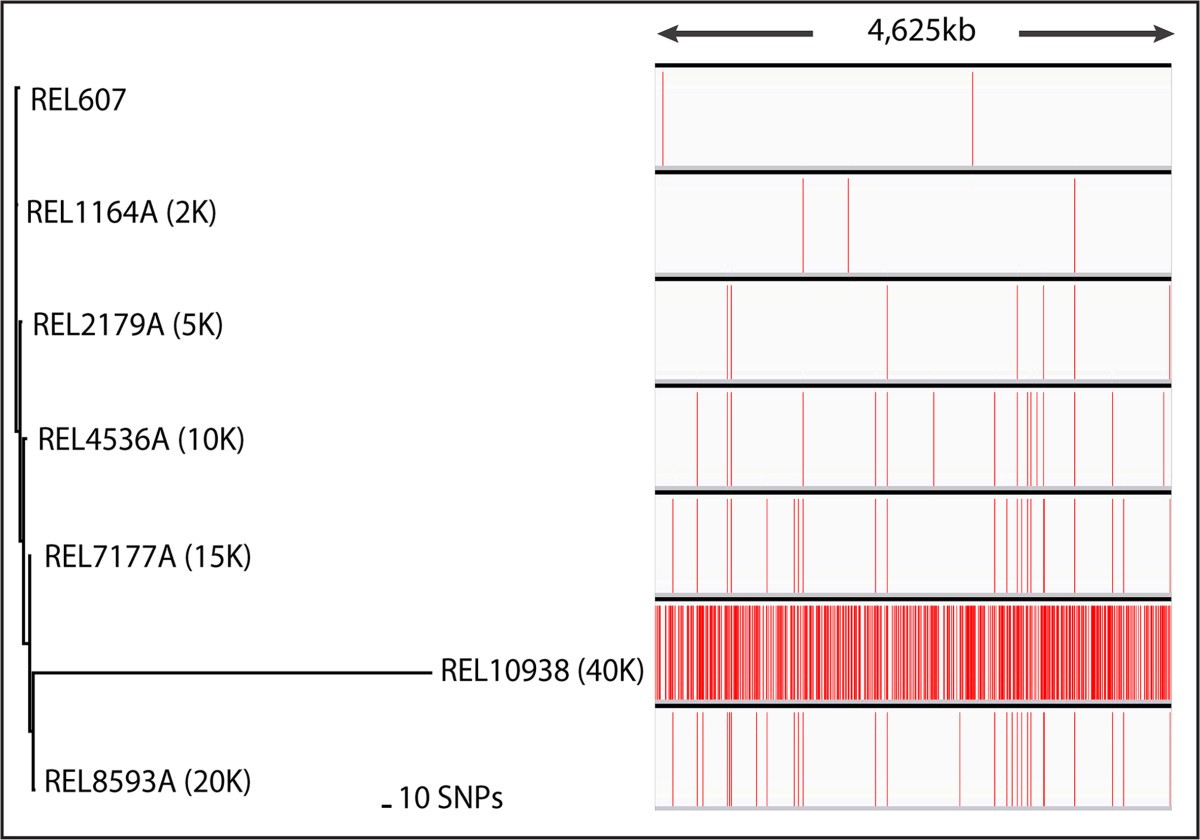
SPANDx: a genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets, BMC Research Notes