dataset - Is there a way to convert smiles format to TUdataset


dataset - Is there a way to convert smiles format to TUdataset format? - Stack Overflow
A Complete Guide to Audio Datasets
sdftosmi.py: Convert multiple ligands/compounds in SDF format to SMILES. — Bioinformatics Review

sdftosmi.py: Convert multiple ligands/compounds in SDF format to SMILES. — Bioinformatics Review
How to convert the molecular graph in SDF/JSON format to the input of graphormer? · Issue #108 · microsoft/Graphormer · GitHub

PDF) Design of a Chemical Graph Database to Query on an Atom-Specific Level

How to convert bundle SMILES to .sdf file format #Molecular_Docking #ChemistryStudio

2103.09430] OGB-LSC: A Large-Scale Challenge for Machine Learning on Graphs
Pytorch-Geometric/pytorch_geometric_introduction.py at master · marcin-laskowski/Pytorch-Geometric · GitHub
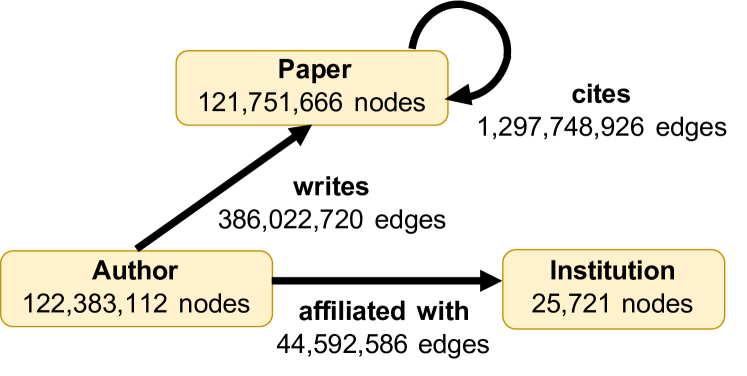
2103.09430] OGB-LSC: A Large-Scale Challenge for Machine Learning on Graphs