GitHub - Rosemeis/emu: EM-PCA for Ultra-low Coverage Sequencing Data
EM-PCA for Ultra-low Coverage Sequencing Data. Contribute to Rosemeis/emu development by creating an account on GitHub.

PDF) Allelic bias when performing in-solution enrichment of ancient human DNA
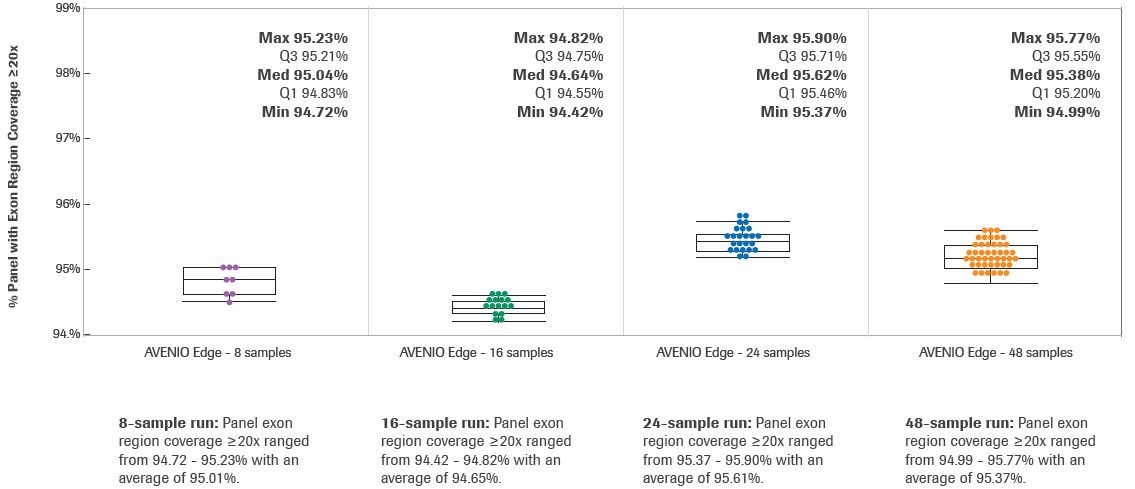
Using HyperExome probes for DNA hybridization workflows

SNPs with highly differentiated allele frequencies with respect to

GitHub - kasumaz/AdultOLgenesis: Single cell analysis of the generation and regulation of oligodendrocyte lineage cells from the adult subventricular zone.

Extensive and accurate benchmarking of DIA acquisition methods and software tools using a complex proteomic standard

PDF) Inferring Population Structure and Admixture Proportions in Low Depth NGS Data
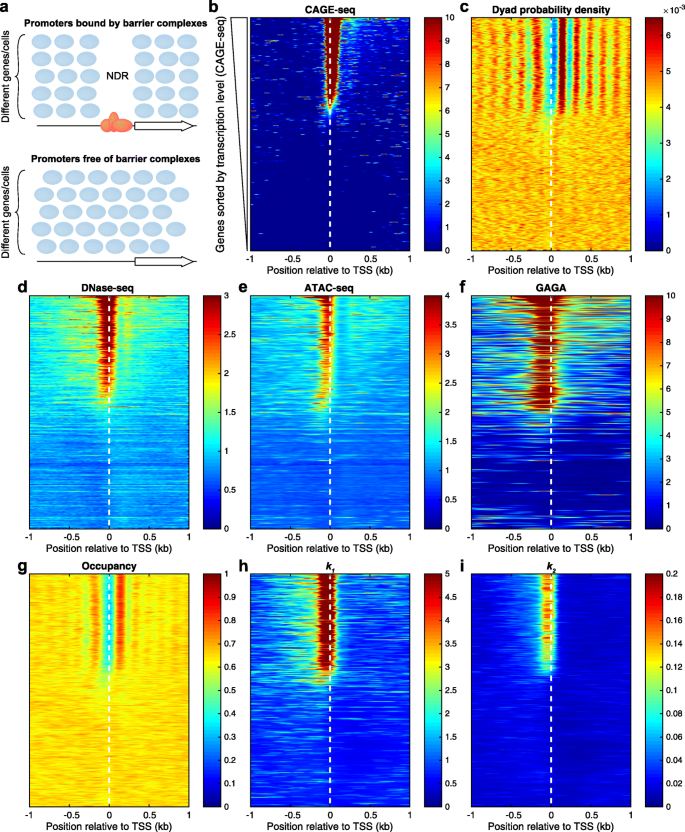
Quantitative MNase-seq accurately maps nucleosome occupancy levels, Genome Biology

A hierarchical clustering of individuals from the HapMap, HGDP and TGP

Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition
KAPA HyperExome - Roche Sequencing Store

PDF) Large-scale Inference of Population Structure in Presence of Missingness using PCA